Our healthcare system, with all its sensitive and interconnected medical information, would be a prime example of the application of digital information systems where tracking each step delivers proof of good quality work. Scientific progress is ongoing, and technologies have been of great support. However, the IT framework in terms of information flow is still lacking, such as the consistent introduction of electronic health records (EHR). A data standard for genomic diagnostics could trailblaze in consolidating the medical data and its metadata (e.g., audit trails of diagnostics) into one place with a clear and easy-to-follow trail for patients and professionals.
Healthcare systems worldwide are grappling with unprecedented challenges, including increased life expectancy, technological advances and a flood of regulations and protocols, as well as shrinking budgets. [1] Especially regulations and protocols – necessary to deliver the highest quality services for patients – rely most of the time on time-consuming documentation. These information trails, particularly in a medical laboratory setting, resulted in a vast amount of manually filled out papers and losing precious time for the actual analysis.
Still, most people will experience at least one diagnostic error in their lives—including errors in lab tests in medical laboratories. Even a tiny error can lead to mistakes like wrongly identifying a patient or mixing up a specimen with potentially negative outcomes.[2] However, as mistakes are human, keeping track of laboratory processes, usually in a well-functioning Quality Management System (QMS) that is mandatory for any medical laboratory, increases the chance of improving the way a laboratory works and minimize the risk for new errors.
In the following, we would like to shine a light on the laboratory workflow in genomics, the information flow along this path and how the MPEG-G standard can set up a framework to function as an intuitive container for all necessary information concerning the trail of quality work in the laboratory.
How is the laboratory workflow in genomics?
In the routine work of a genomics laboratory, multiple steps transform the information in the blood/saliva sample – containing our DNA – into actionable information for potential treatment decisions. This workflow is accompanied by a steady flow of information needed to deliver reliable results and ensure the highest quality level. As genetic testing has become more adapted in the routine diagnostics of patients, the entire process has evolved into a highly automated system including more advanced technical support, for example, pipetting robots.
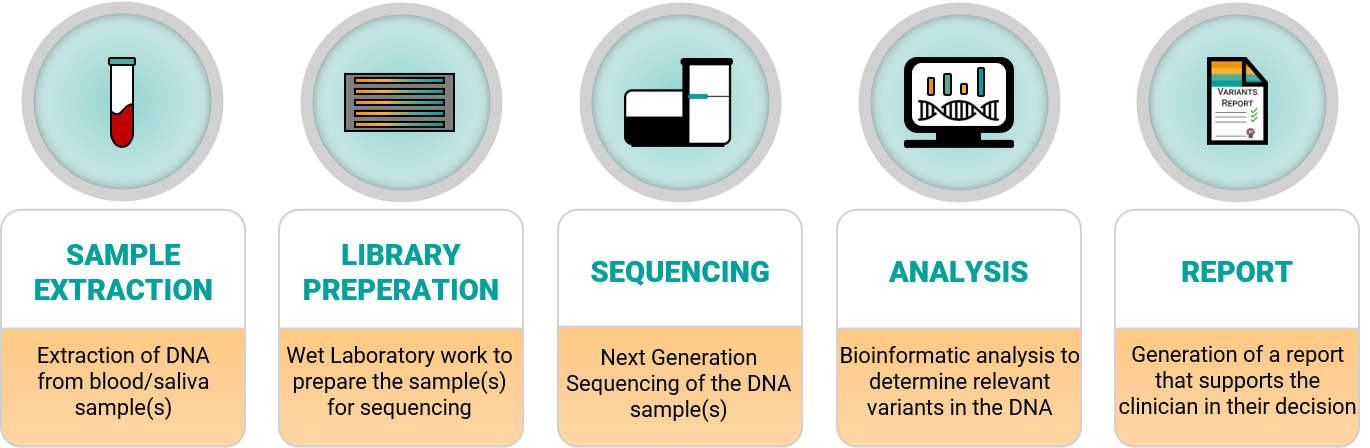
Figure 1 – Laboratory workflow including Sample Extraction, Library preparation, Sequencing, Analysis and Report generation.
What information is generated and what IT systems are currently in place along this process chain?
The flow of information has grown over time as more extensive regions in our DNA were sequenced, and our understanding of how our genome can be responsible for specific diseases increased. The primary driver for this was the evolution from Sanger Sequencing to the Next Generation Sequencing method. The broad scope of what to examine on our DNA and the approach to sequence multiple samples in parallel allowed for the exponential growth of genomic data. The resulting complexity and need for automation are today well-handled inside the wet lab workflow, but interfaces amongst the information flow are still a delicate issue. One of the main issues is the heterogeneity of the multiple software used in the genetic testing workflow. As most of them operate in their own format in terms of Laboratory Information System, the implementation to combine the following data output of the Sequencing and Analysis, that include on their own up to five different legacy formats (FASTQ, SAM, BAM, CRAM and VCF), into a consistent stream of data is time consuming and challenging.
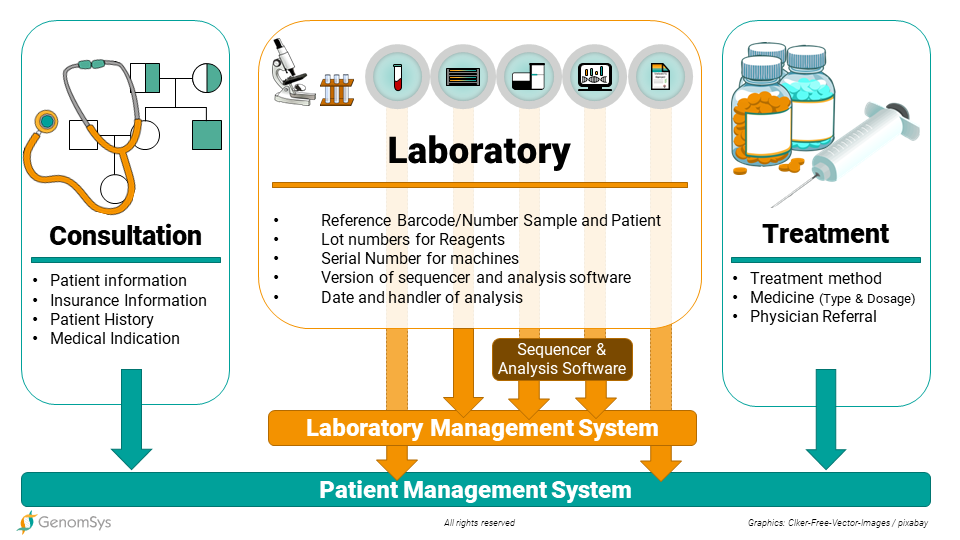
Figure 2 – Data flow in a genomic Laboratory including Patient Management System, Laboratory Information System and Analysis Software.
How can the MPEG-G standard for genomic data help laboratories track their data trail?
The generation of genomic data is a very delicate and complex undergoing. Genomic testing – analyzing our most personal information – has been a key driver of the exponential growth in medical data over the last decade. As Bioinformatician are already successfully handling this pile of Big Data and connecting raw sequencing data with multiple variant annotations, a further connection with other medical data would not be far-fetched.
However the currently used formats have evolved from in-house bioinformatic pipelines without having an additional focus on standardization to provide interoperability in an easy way. This lack of interoperability now leads to an even more complex interaction for including or connecting the precious data with other medical relevant information.
MPEG-G was developed – as is common for any other ISO standard – to ensure a high degree of interoperability in a standardized and easy to follow way. MPEG-G (ISO/IEC 23092) is a series of ISO international standards for the representation of genome sequencing data and associated metadata. This new standard was developed by the Moving Picture Experts Group (MPEG), one of the most prolific ISO working group with more than 1’700 delegates from over 40 countries worldwide. This group has been developing globally used digital media encoding, transmission, and processing standards for more than 30 years. Any individual or organization can join the ISO/MPEG-G working group to keep the spirit for an open standard truly alive.
Besides the efficient capacities of MPEG-G to store and handle genomic data, especially the inclusion of laboratory metadata plays an important role. For example, metadata can be any information along the process chain of the laboratory like Patient history or references to reagents, sequencer or analysis software. So, the MPEG-G file for an individual patient becomes a single consistent file, easy to retrieve.
Additionally, and this is important to automatize the audit trail, MPEG-G allows to include information that are relevant for the laboratory on tracking the data trail when the sample was handled and who extracted, prepared and analyzed it. This time stamp and responsibility tracking, can then be automatized and again stored within a single file to simplify the look up in case it is needed.
Video 1 – MPEG-G operates, as a container of genome sequencing data and associated metadata. The single file for a patient then simplifies keeping track of the data trail for Quality assurance and potential other QMS purposes.
By Lucas Laner on September 7th, 2022.
References:
[1] Prof Trish Greenhalgh and Dr Chrysanthi Papoutsi; Understanding Complexity in Health Systems: International Perspectives (unknown). https://www.biomedcentral.com/collections/complexity
[2] Robert Fenton; The 12 essentials of quality management in laboratory environments (2022). https://www.qualio.com/blog/quality-management-in-laboratory-environments
Picture Source: pexels / pixabay





