Over the past decade, the world has been experiencing a drastic increase in data generation. Since 2013 the global digital data volume has grown by more than 900% and is expected to keep growing further exponentially.[1] In this century, technological breakthroughs have made it possible – especially in the field of genomics – to increase the scope of the spotlight on our DNA as a basis to discover and learn more about ourselves. Researchers, particularly in population genomics, can now compare thousands of genomes amongst a population to understand better the human body and the evolution of our DNA. But the sheer mass of data brings its own challenges. It is time to provide those who work in the spirit of scientific progress with appropriate tools to improve our health continuously.
In the following, we would like to explain further the field of population genomics, the impact the next-generation sequencing (NGS) technology made and will make in this field as well as how a standard designed to truly support the handling of large genomic datasets could be of help for accelerated scientific progress in the decoding of our code of life.
What is Population Genomics?
Population genomics is the large-scale application of genomic technologies to study populations of individuals. It studies heritable features in populations through space and time, setting the ground for understanding evolutionary change.[2] Technically, it examines the causes and ways the frequencies of alleles and genotypes change over time within and between populations. As a result, population genomics allows scientists to understand how ethnic groups are related over time, allowing scientists to understand how our ancestral background has been shaped over time in terms of the environment in which our ancestors have lived their lives.[3] From this, combinations of variations on the DNA can be deduced that may cause disease in one ethnic group while having no effect on health in another. This information helps to make the genetic variation databases even more powerful to increase the significance of genetic testing, taking into account the ethnic origin.
Population genomics has, ever since Darwin, been of interest. However, scientists such as Lewontin and Krakauer (1973) were restricted in the early days due to the sequencing capabilities. They compared multi-locus datasets from multiple populations and identified non-neutral or outlier loci by contrasting patterns of population divergence among genetic regions. All this was done with cumbersome methods that covered only small areas of DNA. But the evolution of genomic testing, due to the transformation to NGS methodologies and greater computational power, allow now to study patterns of genomic divergence on an unprecedented scale.[4]
Today, multiple initiatives around the globe, like the 100’000 genomes project in the United Kingdom, the Turkish Genome Project and Dubai Genomics, aim to collect genomes from their population to gain the beforementioned insights to understand their population’s DNA better. The overall aim is to improve the genomic databases of variations to push forward personalized medicine, through genomics, for better healthcare.
The impact of NGS – delivering the basis of a broader comparison
Since NGS, next-generation sequencing, was introduced in 2005, it allows for parallel sequencing of multiple genes.[5] A quantum leap for genomics to translate biological information into digital for more than just some small areas on the DNA.
The switch from the previous method, Sanger sequencing, which allowed an analysis of a few DNA fragments at a time (length < 1000 base pairs), to the parallel sequencing of multiple genes through next-generation sequencing is why genomics is now facing significantly bigger genomic datasets. As the previous gold standard, Sanger sequencing could only produce a couple of Gigabytes within a couple of months; today, sequencing produces a data flow of multiple Terabytes each week and soon even within days, especially given the capability to sequencing genomes faster. The size of a whole-genome sequencing (WGS) dataset nowadays depends on the format and particular attributes, such as the coverage of the genetic area, the sequencing kit, the sequencing machine, and the number of samples within the sequencing run.
The ever-increasing adoption of NGS-based methods in clinical settings has dramatically advanced the identification of the genetic causes of Mendelian phenotypes today, providing support for diagnostic, preventive, and therapeutic strategies, and supporting the development of personalized medicinal approaches is expected in upcoming years.[6][7] But with the advent of routinely sequencing whole genomes, researchers in population genomics will now be able to compare DNA data on a broader scale.
The foundation is built to receive the entire sequence of DNA. In 2002, almost the entire genome was decoded, missing “only” 8%. But in March this year, scientists achieved the complete decoding of the human genome just perfectly in time for the 20th anniversary of uncovering the sequence of the DNA.[8] Now, it seems to be only a matter of time before enough genomes will be sequenced and compared to each other to discover further information needed during genomic testing. But as in any other industry handling many data, Big Data itself produces challenges. The use of NGS brings concerns and drawbacks associated with the demanding process of storing, processing, analyzing, and interpreting of the massive amount of generated genomics data.4
And although the evolution of sequencing has been very rapid, another bottleneck has emerged that is not in the wet lab but challenges the IT framework. Precious time is spent processing the datasets, ultimately cutting the researchers’ time in their actual work of interpreting genomic data. To compensate for precisely this unnecessary lost time, the Moving Picture Experts Group (MPEG) started back in 2016 developing the first and only open standard specification, published by ISO, for the more efficient compression and transport of next-generation sequencing data.[9]
Particularly for Population Genomics, the MPEG-G standard can deliver benefits in terms of faster transmission, less storage need and faster access to the desired region amongst the DNA sequence or the annotated results.
The compression benefit of MPEG-G
When talking about multiple whole-genome sequencing datasets, the size for such information is, to say the least high. Depending on its coverage depth and compression, a single human WGS can account for about 60 – 70 GB in legacy formats. Now in the case of population genomics, the number of datasets is more in the range of hundreds of thousands to millions.
Here is where one of the main benefits of the ISO/IEC 23092 MPEG-G standard comes into play: the efficient compression for genomic datasets. Thus far, the widely used legacy formats have been developed mainly in a setting of a couple of whole genomes, primarily developed within their own habitat of bioinformatics on an academic level for past research purposes with fewer data and smaller data size.
The MPEG-G standard is built to compress the genomic data more efficiently to provide genomic organizations with a format that allows them to store their data in smaller files resulting in fewer costs and faster transmission in an ever more connected world. So, as the beforementioned example of the 100’000 genomes project, the storage benefit, by using MPEG-G compared to the gzipped FASTQ format for the genomic data would amount to 5’217 Terabytes which in the case of cloud storage with AWS services would amount to monthly cost savings of USD122’600.[10]
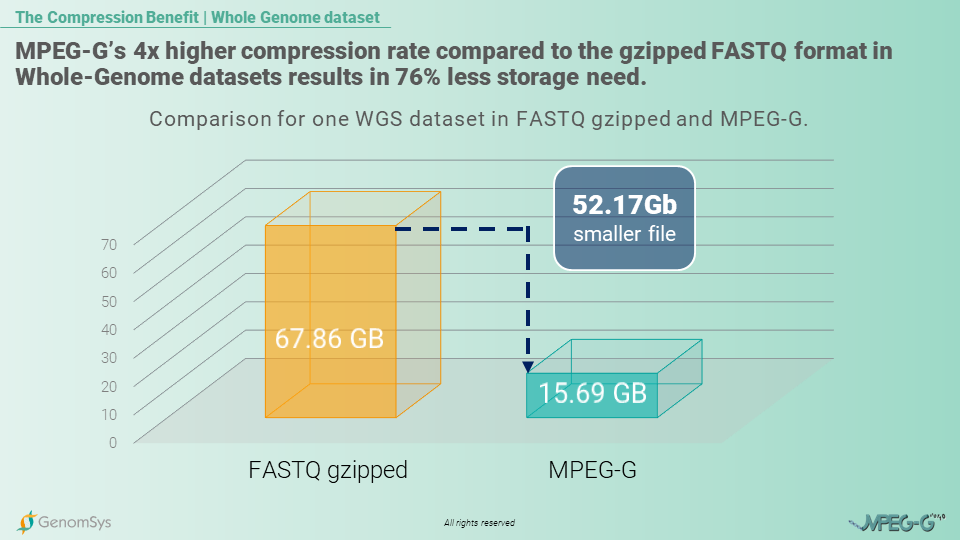
Graphic 1 – Comparison of size for a single WGS genomic datasets in gzipped FASTQ and MPEG-G.[11]
So, smaller files would benefit researchers in Population Genomics to handle the full information of a whole genome but with fewer costs for storage and, overall, faster transmission of the files. This MPEG-G benefit thus makes handling an enormous amount of data – and in population genomics, we talk about thousands of genomes – more efficient in terms of monetary and time resources.
The Selective Access benefit of MPEG-G
Another key advantage is MPEG-G’s Selective Access, with the ability to access data selectively instead of sequentially. This feature is vital for accessing specific regions within the raw DNA sequence – such as chromosomes, genes or even single bases – or scanning for variations in the annotated section. Researchers can then access the data selectively and not sequentially, which is common in legacy formats, and data processing becomes faster. Mainly the time benefit substantially grows with the size of the files.[12] But how is MPEG-G able to access data selectively and not sequentially?
The core of the Selective Access lies within the indexing structure of MPEG-G, which is natively embedded with the data. The indexing step and the sorting step – although sorting in MPEG-G is not strictly required – are inherent parts of the encoding process. Hence they can be performed concurrently with the alignment, for example, without any overhead in terms of processing time: this allows later rapid access to the desired genomic region without the need to use external indexing information to be built before accessing the data.
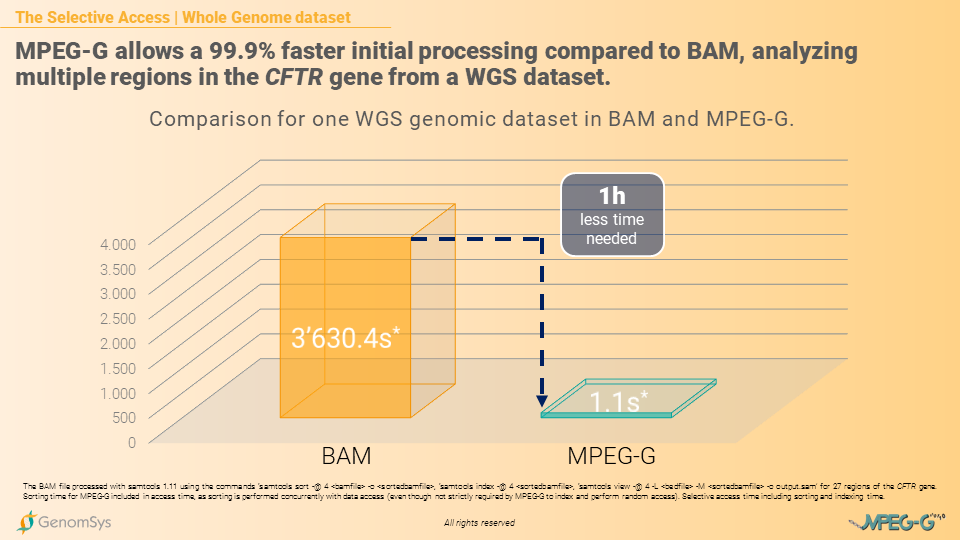
Graphic 2 – Comparison of processing time for WGS genomic datasets in BAM and MPEG-G.[13]
Also, MPEG-G datasets can be transmitted with minimum delay between data generation and consumption thanks to its access unit structure. So each access unit is an independent data chunk and can be as small as needed, making the consumer able to process data as soon as the first access unit is received, thus minimizing the overall data processing time in the genomic pipeline.
This “transport feature”, which is similarly used in streaming videos today by companies like Netflix, avoids being stuck with long waiting times between different pipeline steps and guarantees quicker data consumption by the analysis tools. In legacy formats, usually, the process is sequential from sequencing to variant calling, possibly including streaming over the network. Ultimately consumption of the data, which is in genomics, the analysis of the data, is delayed with regards to a scenario, like the one made possible by MPEG-G, where each step can start without waiting for the previous ones to be completely done. For example, even on a tiny fraction of DNA, the corresponding aligned data can be sent through the network, and researchers in population genomics can immediately start comparing datasets to find similarities amongst vast amounts of data.
Video 1 – MPEG-G’s compression feature and Selective Access can provide an efficient framework for researchers in population genomics handling vast amounts of data. The Selective Access allows for a rapid scan for desired similarities while comparing multiple WGS datasets. The compression helps in terms of storage costs and speed of transmission.
GenomSys professional’s solution leveraging the interoperability feature from MPEG-G
All of our solutions run natively in MPEG-G, making them the ideal entry points for organizations in genomics to take advantage of this new unique and efficient genomic standard.
GenomSys helps organizations handle large amounts of genomic data to store them cost-efficiently and be equipped for the new era of genomics with an ever-increasing volume of data. Via GenomSys MPEG-G Toolkit, a collection of CE-marked software tools to process genomic data compliant MPEG-G, allows for encoding losslessly genomic datasets into the new genomic standard, and organizations can leverage the benefits directly.
Furthermore, the Toolkit comes with an intuitive interface to encode and decode the information as well can be implemented by GenomSys in an automatic process directly connected with the sequencing machine, even further pushing the acceleration of processing time.
By Lucas Laner on November 14th, 2022.
References:
[1] Michael Kroker; Big Data sorgt schon 2016 für Speicher-Engpass; 2020 fehlt Speicher-Volumen von 6 Zetabytes (2015). https://blog.wiwo.de/look-at-it/2015/05/05/big-data-sorgt-schon-2016-fur-speicher-engpass-2020-fehlt-speicher-volumen-von-6-zetabytes/
[2] A. Amorim, Population Genetics, Editor(s): Stanley Maloy, Kelly Hughes, Brenner’s Encyclopedia of Genetics (Second Edition), Academic Press, 2013, Pages 407-411, ISBN 9780080961569, https://doi.org/10.1016/B978-0-12-374984-0.01195-5
[3] Charles Rotimi; POPULATION GENOMICS (2022). https://www.genome.gov/genetics-glossary/Population-Genomics
[4] Nosil, P. & Buerkle, A. (2010) Population Genomics. Nature Education Knowledge 3(10):8
[5] Jamuar SS, Tan EC. Clinical application of next-generation sequencing for Mendelian diseases. Hum Genomics 2015 Jun 16;9:10-015-0031-5.
[6] Rabbani B, Tekin M, Mahdieh N. The promise of whole-exome sequencing in medical genetics. J Hum Genet 2014 Jan;59(1):5-15.
[7] Chong JX, Buckingham KJ, Jhangiani SN, Boehm C, Sobreira N, Smith JD, et al. The Genetic Basis of Mendelian Phenotypes: Discoveries, Challenges, and Opportunities. Am J Hum Genet 2015 Aug 6;97(2):199-215.
[8] Evan Bush; A human genome has finally been fully decoded (2022). https://www.nbcnews.com/science/science-news/human-genome-finally-fully-decoded-rcna22029
[9] Moving Picture Experts Group – Genomics; MPEG-G ISO/IEC 23092 (2022). https://mpeg-g.org/
[10] In-House calculation of the same 100’000 WGS dataset with 30x coverage in each format: FASTQ gzipped 67.86GB; MPEG-G file=15.69G; AWS S3 standard pricing=0.0235USD per GB and month
[11] In-House measurements of the same WGS dataset with 30x coverage in each format: FASTQ gzipped file pairs= 67.86GB; MPEG-G file=15.69G.
[12] National Computer Conference and Exposition (1957)
[13] The BAM file processed with samtools 1.11 using the commands ‘samtools sort -@ 4 <bamfile> -o <sortedbamfile>, ‘samtools index -@ 4 <sortedbamfile>, ‘samtools view -@ 4 -L <bedfile> -M <sortedbamfile> -o output.sam’ for multiple regions of the CFTR gene. Sorting time for MPEG-G included in access time, as sorting is performed concurrently with data access (even though not strictly required by MPEG-G to index and perform random access). Selective access time including sorting and indexing time.





